This is my final submission for GSoC and it contains description of my work and the journey of the project.
I started with translating all the SPM dependencies in the community bonding period itself. SPM dependencies are functions of the SPM library in MATLAB. I was able to translate them with the help of the MATLAB code available for each of them at https://github.com/spm/spm12.
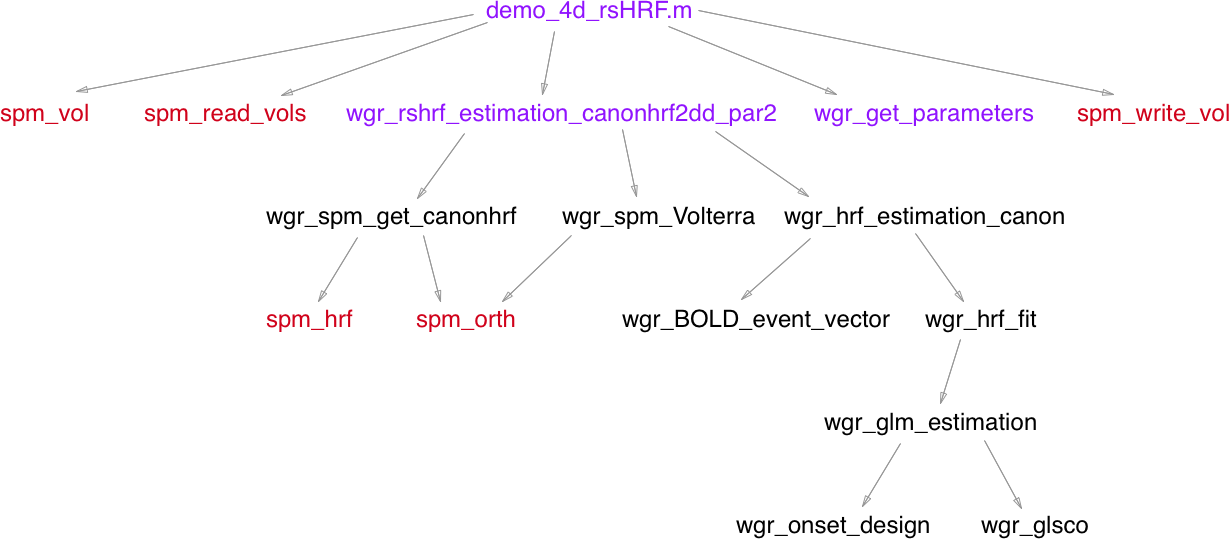
I started with 3 files initially which were demo_4d_rsHRF.m, wgr_rshrf_estimation_canonhrf2dd_par2.m and wgr_get_parameters.m.
Before I began the conversion, I made dependency graphs. These graphs helped me in finding a topological order for translation of functions.
The graph for STAGE-1 is given below:
Issues Faced:
a) MATLAB uses 1 based indexing while Python uses 0 based indexing
b) Everything in MATLAB is a multi-dimensional array
c) Default behaviors of many different mathematical implementations differ
d) Rounding Off Numbers
I was able to work around them through repetitive testing and was able to reproduce results for the first demo.
The other 2 files that were remaining to be translated were demo_main_deconvolution_FIR.m and hrf_retrieval_and_deconvolution_para.m.
The dependency graph for these 2 files and their associated functions is given below:
For the translation part that ended here, I mostly referred to https://docs.scipy.org/doc/numpy/user/numpy-for-matlab-users.html
However, it was required to test each function repeatedly with the sample data to ensure correctness at each step of translation. Guaranteeing this was quite hard and took significant amount of time (partly because of my slow PC).
After fixing all inconsistency issues, I ended up with 3 kinds of demos aka estimation methods that are canon2dd, FIR, sFIR.
With 2 sample datasets available and 3 kinds of analysis (as mentioned above) + 5 output files for each kind of analysis gave a total of 5*3*2 = 30 output files. Running the script with MATLAB to get these 30 files and then with my translated code further helped me debug where the output mismatched between the files.
With the surety that the translated version now worked correctly, the next step was to implement command - line arguments.
First, the design of the command-line arguments was discussed. Specifically, which all arguments to set parameters, which all to take input, and which all to set flags for features. Once this was done, error handling was discussed. The design of the application as a whole required quite a lot of discussion. Especially the errors that the app should handle.
This + error handling was done with the help of the argparse library (https://docs.python.org/3/howto/argparse.html#id1).
Some advanced uses such as getting a dictionary of parameters of a certain group in argeparse was taken from StackOverflow.
Till this step, the application worked as a normal CLI should work, but work was still left so that it could be converted to follow the specifications of how a BIDS - App should work.
The application was supposed to work on BIDS - Datasets. These datasets were to be queried over using the PyBIDS package.
The tutorial over here helped a lot with this : https://miykael.github.io/nipype_tutorial/notebooks/basic_data_input_bids.html
Next, I worked on renaming all files and organised them into 4 sub-packages along with 1 main package. The tutorial over here helped me with resolving complex interleaved imports: https://docs.python.org/3/tutorial/modules.html.
The next step was to release the application for users to test out. For this, the package we developed was to be released to the
internet so that they could install it with just one command pip3 install rsHRF. The tutorial present here helped me with
writing the setup.py file: https://packaging.python.org/tutorials/packaging-projects/
The app used to analyse all the subjects in the dataset (queried with PyBIDS in Phase - 2). The ability to choose only some subjects out of all available was a BIDS - Specification. This was another task that was completed.
The app was supposed to be wrapped up in a Docker Image with the entrypoint being our CLI tool, installable with pip.
This task introduced me to Dockers. It turned out that there was no restriction on which Base Image to choose to build our app.
Naturally, I first chose a Python:3.6 image which resulted in a very minimal Dockerfile having just 3 lines of code. This was simple,
but resulted in an image size of around 1.35 GB. This was a bit ridiculous for an app that small. I started discovering methods to reduce
the resulting Image size and was able to reduce it to 876 MB. On digging a bit further, I was able to reduce the whole size to 204 MBs with
the compressed version being just about 69 MB. I was finally able to come up with an optimized Dockerfile that also worked.
This process taught me about the concepts of layers in Dockers and how it was useful to remove build files after installation in the same layer
to prevent aggregation in size.
The following Dockerfile was used as a reference: https://github.com/frol/docker-alpine-python-machinelearning/blob/master/Dockerfile
The MATLAB code used parfor (a parallel for loop) in 2 of its functions. The next part of this project was to do something similar so as to achieve gains in speed. To implement this, I started researching about various libraries that can help in implementing this. During this phase, I got to learn about the limitations of Python in multi-threading (due to the python interpreter having something known as a GIL [Global Interpreter Lock]). Thus, multi-processing was the only way to go. This was easily implementable. The next level however, that is, to analyse subjects in parallel gave us troubles. Implementing analysis of subjects in parallel which further called functions having the parfor capability was a case of nested parallelism. While I tried to implement this, it turns out that it is nearly impossible to do so. I did end up exploring many libraries in the process though - Dask, joblib, multiprocessing, etc. I was able to find a hacky solution at https://stackoverflow.com/questions/32688946/create-child-processes-inside-a-child-process-with-python-multiprocessing-failed/32689245. Implementing this resulted in negligible improvement. Thus, we ended up dropping the Idea together since the users could still send subjects to nodes on a high performance cluster manually.
Finally, we ended up integrating CircleCI into our github repository. This was a totally new experience for me. CircleCI enabled us to release new versions for our application by just using git tags. Sending a pull request with a git tag enabled circleCI to automatically deploy the new version of the package at PyPI as well as an updated docker image to DockerHub. Resource used: https://circleci.com/docs/2.0/building-docker-images/
Finally, I wrapped it up with some documentation using GitHub Pages.
Note : The original repository was moved to the BIDS-Apps organisation [https://github.com/BIDS-Apps/rsHRF]. Thus, the merged PRs are present in the old one. [https://github.com/compneuro-da/rsHRF]
Repository : https://github.com/BIDS-Apps/rsHRF
Commits : https://github.com/BIDS-Apps/rsHRF/commits/master
Merged PRs : https://github.com/compneuro-da/rsHRF/pulls?q=is%3Apr+is%3Aclosed+author%3Amadhur-tandon
Documentation : http://bids-apps.neuroimaging.io/rsHRF/
PyPI package : https://pypi.org/project/rsHRF/
BIDS-App Docker Image : https://hub.docker.com/r/bids/rshrf/
-
Group Level Analysis
-
Scripts to help users run the app on HPCs