Last active
August 4, 2020 07:28
-
-
Save maedoc/b377fb8b80c3217222d5774ba47dcce3 to your computer and use it in GitHub Desktop.
SDDE estimation with Stan
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| import pystan | |
| try: | |
| model # avoid recompiling during IPython/notebook session | |
| except: | |
| model = pystan.StanModel('sdde.stan') | |
| data = dict( | |
| nt=500, | |
| d=50, | |
| dt=0.1, | |
| I=0.1234, | |
| K=-1.0, | |
| eta_sd=0.5, | |
| obs_sd=0.5, | |
| ) | |
| run = model.sampling(data=data, iter=500, warmup=250) | |
| samp = run.extract() | |
| xth = samp['x_t_hat'].mean(axis=0) | |
| figure(figsize=(15, 15)) | |
| subplot(211); | |
| plot(xth); | |
| title('X(t) inferred') | |
| subplot(223); hist(samp['K_hat'], 100); title('True K=-1.0'); | |
| subplot(224); hist(samp['I_hat'], 100); title('True I=0.1234') |
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| data { | |
| int nt; // num time points | |
| int d; // delay in steps | |
| real dt; // time step size | |
| real I; // vertical shift bistable system | |
| real K; // scale of delay input | |
| real eta_sd; // scale of SDE noise | |
| real obs_sd; // scale of observation noise | |
| } | |
| transformed data { | |
| vector[nt] x_t; | |
| vector[nt] eta; // noise | |
| for (t in 1:nt) | |
| eta[t] = normal_rng(0, 1); | |
| for (t in 1:d) | |
| x_t[t] = eta[t]; | |
| for (t in (d + 1):nt) | |
| { | |
| real x1 = x_t[t - 1]; | |
| real xd = x_t[t - d]; | |
| real dx = x1 - x1*x1*x1/3.0 + I + K * xd + eta[t]*eta_sd; | |
| x_t[t] = x1 + dt * dx; | |
| } | |
| } | |
| parameters { | |
| vector[nt] eta_hat; | |
| real I_hat; | |
| real K_hat; | |
| } | |
| transformed parameters { | |
| vector[nt] x_t_hat; | |
| for (t in 1:d) | |
| x_t_hat[t] = eta_hat[t]; | |
| for (t in (d + 1):nt) | |
| { | |
| real x1 = x_t_hat[t - 1]; | |
| real xd = x_t_hat[t - d]; | |
| real dx = x1 - x1*x1*x1/3.0 + I_hat + K_hat * xd + eta_hat[t]*eta_sd; | |
| x_t_hat[t] = x1 + dt * dx; | |
| } | |
| } | |
| model { | |
| eta_hat ~ std_normal(); | |
| I_hat ~ std_normal(); | |
| K_hat ~ std_normal(); | |
| x_t ~ normal(x_t_hat, obs_sd); | |
| } |
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| import numpy as np | |
| import pylab as pl | |
| import pystan | |
| try: | |
| model | |
| except: | |
| model = pystan.StanModel('sdde2.stan') | |
| dt=0.2 | |
| data = dict( | |
| nt=250, | |
| nh=50, | |
| dmu=50*dt, | |
| dsd=10*dt, | |
| dt=dt, | |
| I=0.1234, | |
| K=-0.5, | |
| eta_sd=0.5, | |
| obs_sd=0.5, | |
| debug=0, | |
| ) | |
| run = model.sampling(data=data, iter=2000, warmup=1000, chains=4) | |
| samp = run.extract() | |
| pl.figure() | |
| pl.subplot(211); | |
| pl.plot(samp['x_t_hat'].mean(axis=0)); | |
| pl.title('X(t) inferred') | |
| pl.subplot(234); pl.hist(samp['K_hat'], 100); pl.title(f'True K={data["K"]}'); | |
| pl.subplot(235); pl.hist(samp['I_hat'], 100); pl.title(f'True I={data["I"]}') | |
| pl.subplot(236); pl.hist(samp['dhat'], 100); pl.title(f'True d={data["dmu"]}') |
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| data { | |
| int nt; // num time points | |
| int nh; // num history for t<0 | |
| real dmu; // delay prior mean | |
| real dsd; // delay prior std | |
| real dt; // time step size | |
| real I; // vertical shift bistable system | |
| real K; // scale of delay input | |
| real eta_sd; // scale of SDE noise | |
| real obs_sd; // scale of observation noise | |
| } | |
| transformed data { | |
| vector[nt] x_t = rep_vector(0.0, nt); | |
| vector[nt] eta; // noise | |
| vector[nt] ts; // times | |
| real w_norm = 1.0 / (dsd * sqrt(2.0 * pi())); | |
| for (t in 1:nt) | |
| { | |
| eta[t] = normal_rng(0, 1); | |
| ts[t] = dt * t; | |
| } | |
| x_t[1:nh] = eta[1:nh]; | |
| for (t in (nh + 1):nt) | |
| { | |
| real x1 = x_t[t - 1]; | |
| vector[nt] wt = w_norm * exp(-0.5 * square((ts - (t*dt - dmu))/dsd)); | |
| real dx = x1 - x1*x1*x1/3.0 + I + K * (wt' * x_t) + eta[t]*eta_sd; | |
| x_t[t] = x1 + dt * dx; | |
| } | |
| } | |
| parameters { | |
| vector[nt] eta_hat; | |
| real I_hat; | |
| real K_hat; | |
| real<lower=0> dhat; | |
| } | |
| transformed parameters { | |
| vector[nt] x_t_hat = eta_hat; | |
| for (t in (nh + 1):nt) | |
| { | |
| real x1 = x_t_hat[t - 1]; | |
| vector[nt] wt = w_norm * exp(-0.5 * square((ts - (t*dt - dhat))/dsd)); | |
| real dx = x1 - x1*x1*x1/3.0 + I_hat + K_hat * (wt' * x_t_hat) + eta_hat[t]*eta_sd; | |
| x_t_hat[t] = x1 + dt * dx; | |
| } | |
| } | |
| model { | |
| eta_hat ~ std_normal(); | |
| I_hat ~ std_normal(); | |
| K_hat ~ std_normal(); | |
| dhat ~ std_normal(); | |
| x_t ~ normal(x_t_hat, obs_sd); | |
| } |
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| dt=0.1 | |
| nt=500 | |
| nh=100 | |
| dmu=100*dt | |
| dsd=30*dt | |
| dt=dt | |
| I=0.1234 | |
| K=-0.5 | |
| eta_sd=0.5 | |
| debug=0 | |
| x_t = zeros(nt) | |
| eta = zeros(nt) | |
| ts = zeros(nt) | |
| w_norm = 1.0 / (dsd * sqrt(2.0 * pi)) | |
| sqrt_dt = sqrt(dt) | |
| for t=1:nt | |
| eta[t] = randn() | |
| ts[t] = dt * t | |
| end | |
| function square(x) x * x end | |
| x_t[1:nh] = eta[1:nh] | |
| for t=(nh + 1):nt | |
| x1 = x_t[t - 1] | |
| wt = @. w_norm * exp(-0.5 * square((ts - (t*dt - dmu)) / dsd)) | |
| dx = x1 - x1*x1*x1 / 3.0 + I + K * (wt' * x_t) | |
| x_t[t] = x1 + dt * dx + sqrt_dt * eta[t] * eta_sd | |
| end | |
| using Turing, Plots, StatsPlots | |
| @model sdde3(dsd, dmu, x_t, w_norm, ts, dt, eta_sd) = begin | |
| I_hat ~ Normal(0, 1) | |
| K_hat ~ Normal(0, 1) | |
| dhat ~ Normal(0, 1) | |
| dhat_ = dhat / dsd + dmu + dsd; | |
| for t=(nh + 1):nt | |
| x1 = x_t[t - 1] | |
| wt = @. w_norm * exp(-0.5 * square((ts - (t*dt - dhat_)) / dsd)) | |
| dx = x1 - x1*x1*x1 / 3.0 + I_hat + K_hat * (wt' * x_t) | |
| # @show x1, dt, dx | |
| x_t[t] ~ Normal(x1 + dt * dx, eta_sd::Float64) | |
| end | |
| end | |
| run = sample(sdde3(dsd, dmu, x_t, w_norm, ts, dt, eta_sd), NUTS(), 1000) | |
| @show describe(run) | |
| @show I, K, dmu | |
| plot(run) |
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| import numpy as np | |
| import pylab as pl | |
| import pystan | |
| try: | |
| model | |
| except: | |
| model = pystan.StanModel('sdde3.stan') | |
| dt=0.1 | |
| data = dict( | |
| nt=500, | |
| nh=100, | |
| dmu=100*dt, | |
| dsd=30*dt, | |
| dt=dt, | |
| I=0.1234, | |
| K=-0.5, | |
| eta_sd=0.5, | |
| debug=0, | |
| ) | |
| run = model.sampling(data=data, iter=2000, warmup=1000, chains=4) | |
| samp = run.extract() | |
| pl.figure() | |
| pl.subplot(211); | |
| pl.plot(samp['x_t_'].mean(axis=0)); | |
| pl.title('X(t)') | |
| pl.subplot(234); pl.hist(samp['K_hat'], 100); pl.title(f'True K={data["K"]}'); | |
| pl.subplot(235); pl.hist(samp['I_hat'], 100); pl.title(f'True I={data["I"]}') | |
| pl.subplot(236); pl.hist(samp['dhat_'], 100); pl.title(f'True d={data["dmu"]}') |
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| data { | |
| int nt; // num time points | |
| int nh; // num history for t<0 | |
| real dmu; // delay prior mean | |
| real dsd; // delay prior std | |
| real dt; // time step size | |
| real I; // vertical shift bistable system | |
| real K; // scale of delay input | |
| real eta_sd; // scale of SDE noise | |
| } | |
| transformed data { | |
| vector[nt] x_t = rep_vector(0.0, nt); | |
| vector[nt] eta; // noise | |
| vector[nt] ts; // times | |
| real w_norm = 1.0 / (dsd * sqrt(2.0 * pi())); | |
| real sqrt_dt = sqrt(dt); | |
| for (t in 1:nt) | |
| { | |
| eta[t] = normal_rng(0, 1); | |
| ts[t] = dt * t; | |
| } | |
| x_t[1:nh] = eta[1:nh]; | |
| for (t in (nh + 1):nt) | |
| { | |
| real x1 = x_t[t - 1]; | |
| vector[nt] wt = w_norm * exp(-0.5 * square((ts - (t*dt - dmu))/dsd)); | |
| real dx = x1 - x1*x1*x1/3.0 + I + K * (wt' * x_t); | |
| x_t[t] = x1 + dt * dx + sqrt_dt*eta[t]*eta_sd; | |
| } | |
| } | |
| parameters { | |
| real I_hat; | |
| real K_hat; | |
| real dhat; | |
| } | |
| transformed parameters { | |
| // shift to have slightly wrong prior | |
| real dhat_ = dhat/dsd + dmu + dsd; | |
| } | |
| model { | |
| I_hat ~ std_normal(); | |
| K_hat ~ std_normal(); | |
| dhat ~ std_normal(); | |
| for (t in (nh + 1):nt) | |
| { | |
| real x1 = x_t[t - 1]; | |
| vector[nt] wt = w_norm * exp(-0.5 * square((ts - (t*dt - dhat_))/dsd)); | |
| real dx = x1 - x1*x1*x1/3.0 + I_hat + K_hat * (wt' * x_t); | |
| x_t[t] ~ normal(x1 + dt * dx, eta_sd); | |
| } | |
| } | |
| generated quantities { | |
| vector[nt] x_t_ = x_t; // too lazy to recode sim in Python! | |
| } |
Author
maedoc
commented
Dec 19, 2019
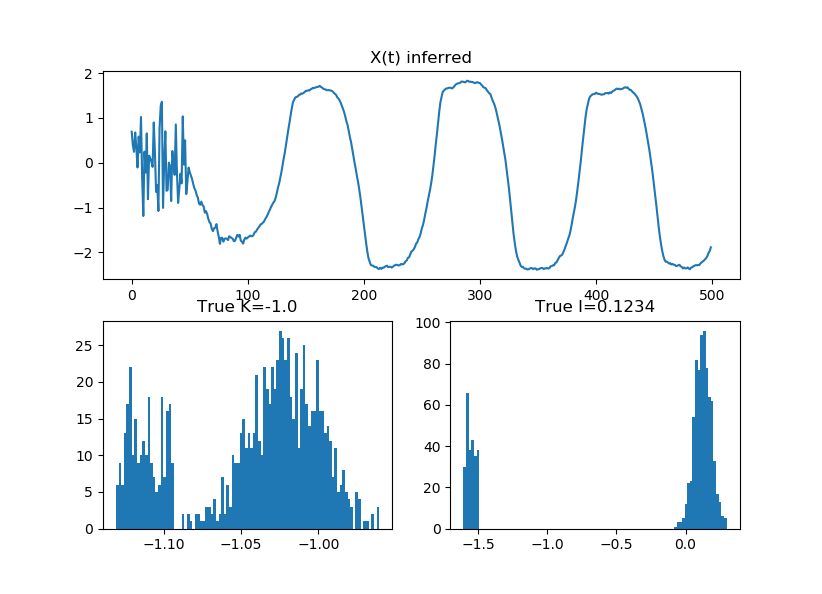
Added second variant for estimating delay but doesn't initialize properly (yet)
Fixed the temporal kernel in the second variant, so it runs now. It's significantly more expensive 💻 🛫 than the first one, but it estimates delay.
Added 3rd variant which doesn't have an observation model or estimate noise, and it's considerably faster. This suggests the 2nd variant could be better constrained to behave nicely.
Added a Julia version with Turing.jl which sort of works but unlikely to be optimal.
Sign up for free
to join this conversation on GitHub.
Already have an account?
Sign in to comment
