Last active
February 15, 2023 07:28
-
-
Save metamaden/810298c5c875905169216ae76ebc1cfd to your computer and use it in GitHub Desktop.
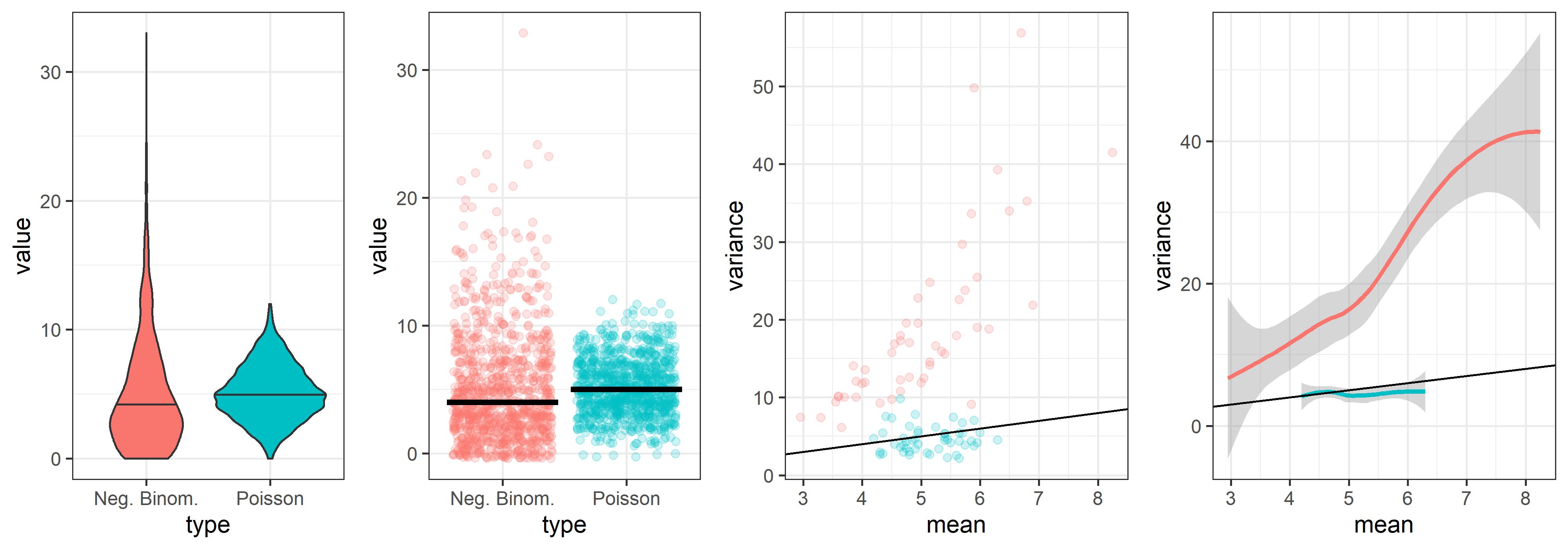
Simulate and plot Poisson and Negative Binomial distributions in R
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| #!/usr/bin/env R | |
| # Author: Sean Maden | |
| # | |
| # Simulate Poisson and Negative Binomial distributions, plotting the point | |
| # values and mean-variance relationships. | |
| # | |
| # load dependencies | |
| libv <- c("ggplot2", "gridExtra") | |
| sapply(libv, library, character.only = TRUE) | |
| # set plot parameters | |
| pt.alpha <- 0.2 | |
| jpg.width = 10 | |
| jpg.height = 3.5 | |
| num <- 5 # layout scale value | |
| jpg.fname <- "dist-examples.jpeg" | |
| # set simulation parameters | |
| seed.num <- 0 | |
| num.points <- 1e3 # total data points | |
| num.groups <- 50 # analogous to genes | |
| set.seed(seed.num) | |
| # simulate new data | |
| set.seed(seed.num) | |
| rn <- rnbinom(n = num.points, size = 2, mu = 5) | |
| rp <- rpois(num.points, lambda = 5) | |
| mrn <- matrix(rn, nrow = num.groups) | |
| mrp <- matrix(rp, nrow = num.groups) | |
| # get violin plots | |
| dfp1 <- data.frame(value = rn); dfp1$type <- "Neg. Binom." | |
| dfp2 <- data.frame(value = rp); dfp2$type <- "Poisson" | |
| dfp.pt <- rbind(dfp1, dfp2) | |
| plot1 <- ggplot(dfp.pt, aes(x = type, y = value, fill = type)) + | |
| geom_violin(draw_quantiles = 0.5) + theme_bw() + | |
| theme(legend.position = "none") | |
| # get jitter plots | |
| dfp1 <- data.frame(value = rn); dfp1$type <- "Neg. Binom." | |
| dfp2 <- data.frame(value = rp); dfp2$type <- "Poisson" | |
| dfp.pt <- rbind(dfp1, dfp2) | |
| plot2 <- ggplot(dfp.pt, aes(x = type, y = value, color = type)) + | |
| geom_jitter(alpha = pt.alpha) + theme_bw() + | |
| theme(legend.position = "none") + | |
| stat_summary(fun = "median", color = "black", geom = "crossbar") | |
| # get mean-variance plots | |
| dfp1 <- data.frame(mean = rowMeans(mrn), | |
| variance = rowVars(mrn)) | |
| dfp1$type <- "Neg. Binom." | |
| dfp2 <- data.frame(mean = rowMeans(mrp), | |
| variance = rowVars(mrp)) | |
| dfp2$type <- "Poisson" | |
| dfp.mv <- rbind(dfp1, dfp2) | |
| plot3 <- ggplot(dfp.mv, aes(x = mean, y = variance, group = type, color = type)) + | |
| geom_point(alpha = pt.alpha) + geom_abline(slope = 1, intercept = 0) + | |
| theme_bw() + theme(legend.position = "none") | |
| plot4 <- ggplot(dfp.mv, aes(x = mean, y = variance, group = type, color = type)) + | |
| geom_smooth() + geom_abline(slope = 1, intercept = 0) + | |
| theme_bw() + theme(legend.position = "none") | |
| # save composite plots | |
| lm <- matrix(c(rep(1, num), rep(2, num), | |
| rep(3, num+1), rep(4, num+1)), nrow = 1) | |
| jpeg(jpg.fname, width = jpg.width, height = jpg.height, | |
| units = "in", res = 400) | |
| grid.arrange(plot1, plot2, plot3, plot4, nrow = 1, layout_matrix = lm) | |
| dev.off() |
Sign up for free
to join this conversation on GitHub.
Already have an account?
Sign in to comment
Plot output