-
-
Save moble/e2b3e89fc0220e4363765fadc8c1dd73 to your computer and use it in GitHub Desktop.
| #!/usr/bin/env python | |
| # Copyright (c) 2020, Michael Boyle | |
| # | |
| # MIT License | |
| # | |
| # Permission is hereby granted, free of charge, to any person obtaining a copy | |
| # of this software and associated documentation files (the "Software"), to deal | |
| # in the Software without restriction, including without limitation the rights | |
| # to use, copy, modify, merge, publish, distribute, sublicense, and/or sell | |
| # copies of the Software, and to permit persons to whom the Software is | |
| # furnished to do so, subject to the following conditions: | |
| # | |
| # The above copyright notice and this permission notice shall be included in | |
| # all copies or substantial portions of the Software. | |
| # | |
| # THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR | |
| # IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, | |
| # FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE | |
| # AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER | |
| # LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, | |
| # OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE | |
| # SOFTWARE. | |
| """Ensure GTF-format files have `gene_id` fields unique to each `gene_name` or `transcript_id`. | |
| You will need numpy and pandas (and optionally tqdm) installed. | |
| Simply call this script as | |
| python unique_gene_id.py file1.gtf | |
| where `file1.gtf` is a file in GTF format, described at https://mblab.wustl.edu/GTF22.html. | |
| Multiple files may be specified in the same command, and will be transformed in turn. Each | |
| new file is output alongside the original, with '.unique_gene_id' inserted before the | |
| original file ending (which is presumably '.gtf'). | |
| """ | |
| try: | |
| from tqdm.auto import tqdm | |
| except: | |
| def tqdm(iterator, **kwargs): | |
| return iterator | |
| def read_gtf(file_name): | |
| """Read GTF file into pandas DataFrame | |
| The GTF format is specified at https://mblab.wustl.edu/GTF22.html. | |
| Note that the "attributes" column is returned as a series of strings. To | |
| convert to `dict` types, call the `convert_attributes` function. | |
| """ | |
| import numpy as np | |
| import pandas as pd | |
| seqname = "string" | |
| source = "string" | |
| features = pd.CategoricalDtype({"CDS", "start_codon", "stop_codon", "5UTR", "3UTR", "inter", "inter_CNS", "intron_CNS", "exon", "transcript"}) | |
| start = int | |
| end = int | |
| score = "string" | |
| strand = pd.CategoricalDtype({"+", "-", "."}) | |
| frame = pd.CategoricalDtype({0, 1, 2, "."}) | |
| attributes = np.dtype("O") | |
| comments = "string" | |
| gtf_column_names = ["seqname", "source", "feature", "start", "end", "score", "strand", "frame", "attributes"]#, "comments"] | |
| gtf_dtypes = [seqname, source, features, start, end, score, strand, frame, attributes]#, comments] | |
| return pd.read_csv(file_name, sep="\t", comment="#", names=gtf_column_names, dtype={k:v for k, v in zip(gtf_column_names, gtf_dtypes)}) | |
| def convert_attributes_to_dict(df): | |
| """Convert GTF DataFrame "attributes" from strings into dicts | |
| Note that this operates in place, meaning that the input DataFrame is changed. | |
| """ | |
| for index, value in tqdm(df["attributes"].iteritems(), total=len(df), dynamic_ncols=True): | |
| if not isinstance(value, str): | |
| continue | |
| attributes = { # Start with required defaults; likely overwritten below | |
| "gene_id": "", | |
| "transcript_id": "" | |
| } | |
| for attribute in value.strip().split(";"): | |
| attribute = attribute.strip() | |
| if attribute: | |
| split = attribute.split(" ") | |
| if split: | |
| if len(split) != 2: | |
| raise ValueError(f"Bad split '{split}' in '{attribute}' of '{value}'") | |
| k, v = split | |
| attributes[k] = v | |
| else: | |
| raise ValueError(f"No split in '{attribute}' of '{value}'") | |
| df.at[index, "attributes"] = attributes | |
| return df | |
| def make_gene_id_unique(df): | |
| """Append suffixes to ensure that `gene_id` is unique | |
| This function searches the input for rows with the same `gene_id` field, but | |
| different values in `gene_name` (or, if that doesn't exist, in | |
| `transcript_id`). If multiple such rows are found, they are modified by | |
| appending a suffix (a "." followed by an integer), to ensure that no such | |
| clashes exist on output. | |
| Note that this operates in place, meaning that the input DataFrame is changed. | |
| """ | |
| from collections import defaultdict | |
| dd = defaultdict(set) | |
| for d in df["attributes"]: | |
| if d["gene_id"]: | |
| dd[d["gene_id"]].add(d.get("gene_name", d["transcript_id"])) | |
| suffix_map = {(k+s): i for k, v in dd.items() for i, s in enumerate(sorted(v)) if len(v) > 1} | |
| for d in df["attributes"]: | |
| if d["gene_id"]: | |
| key = d["gene_id"] + d.get("gene_name", d["transcript_id"]) | |
| if key in suffix_map: | |
| d["gene_id"] = f'"{d["gene_id"][1:-1]}.{suffix_map[key]}"' | |
| def convert_attributes_to_string(df): | |
| """Convert GTF DataFrame "attributes" from dicts into strings | |
| Note that this operates in place, meaning that the input DataFrame is changed. | |
| """ | |
| for index, value in tqdm(df["attributes"].iteritems(), total=len(df), dynamic_ncols=True): | |
| if not isinstance(value, dict): | |
| continue | |
| df.at[index, "attributes"] = "; ".join(f"{k} {v}" for k, v in value.items()) | |
| return df | |
| def unique_gene_id(file_name): | |
| """Rewrite GTF file to have `gene_id` fields unique for each `gene_name` or `transcript_id` | |
| The input `file_name` is read and processed, and a new file is output alongside | |
| it, where ".unique_gene_id" is inserted before the file ending (presumably | |
| ".gtf"). | |
| """ | |
| import csv | |
| import pathlib | |
| path_in = pathlib.Path(file_name).expanduser().resolve() | |
| path_out = path_in.with_suffix(".unique_gene_id" + path_in.suffix) | |
| df = read_gtf(file_name) | |
| convert_attributes_to_dict(df) | |
| make_gene_id_unique(df) | |
| convert_attributes_to_string(df) | |
| with path_out.open(mode="w", newline="") as f_out: | |
| with path_in.open(mode="r") as f_in: | |
| while True: | |
| line = f_in.readline() | |
| if line.startswith("#"): | |
| f_out.write(line) | |
| else: | |
| break | |
| df.to_csv(f_out, sep="\t", header=False, index=False, quoting=csv.QUOTE_NONE) | |
| return df | |
| if __name__ == "__main__": | |
| import sys | |
| if len(sys.argv) < 2 or sys.argv[1] in ["-h", "--help"]: | |
| print( | |
| "Pass one or more GTF-format file names to this script to make all\n" | |
| "`gene_id` fields unique to each `gene_name` or `transcript_id`. The\n" | |
| "input file is read and processed, and a new file is output alongside\n" | |
| "it, where '.unique_gene_id' is inserted before the file ending (which\n" | |
| "is presumably '.gtf')." | |
| ) | |
| else: | |
| for file_name in sys.argv[1:]: | |
| print(f"Converting {file_name}") | |
| unique_gene_id(file_name) |
This was actually a script I put together for my wife; I don't actually work with these files. So I apologize if I misremember the details.
It looks like the problem we had was that some entries were missing the gene_name field, and only in those cases did we use transcript_id as a fallback. So if you're sure that every entry has a gene_name, I don't think the transcript_id will even come into it. You can be sure of that by just removing the line
"transcript_id": ""and then changing every occurrence of
d.get("gene_name", d["transcript_id"])to
d.get("gene_name")With those changes, the code won't even look at transcript_id, so it really shouldn't come into it.
On the other hand, if you're not sure that every entry has a gene_name, I don't know how you'll be able to create a unique gene_id at all for those entries. If you just want to skip them, you can put this in front of those d.get lines:
if not d.get("gene_name", ""):
continueHope that helps.
The second option got me to where I wanted to be, thank you so much for your help!
The second option got me to where I wanted to be, thank you so much for your help!
Hi, there
I have used the stringtie to merge two files wiout -G annotaion.gff3. I just merged the two files : stringtie --merge -o merged.gtf ./EVM.all.gff ./genes_liftoff_longest.gff3 and I obtained the merged.gtf file (seen following pictures)
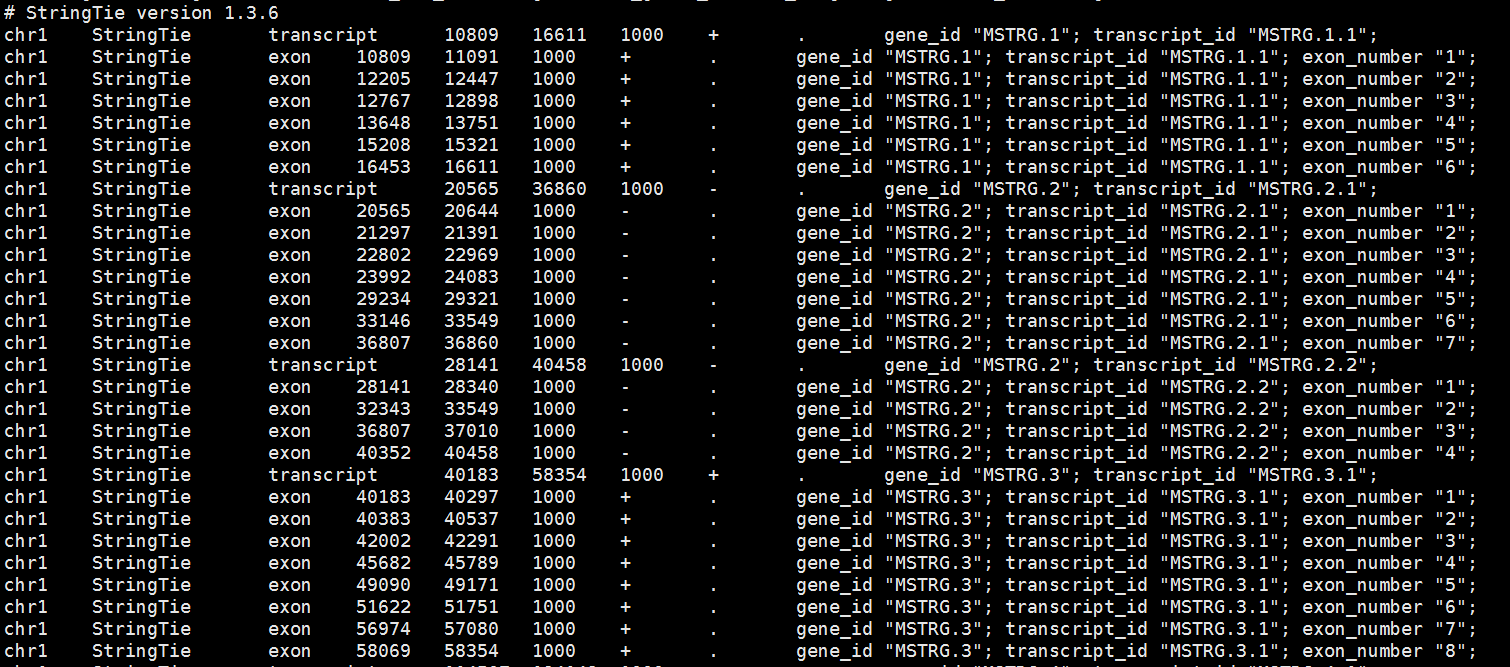
I want to know how to use this scripts? Should I blast the cds squences to annotate the merged.gtf and then use this script ?
Thanks
Sincerely
Yizhong Huang
Hello, I just tried using the script and it worked great, but I'm not interested in differentiating transcripts. Is there a way to modify this to only create unique gene_id names for those that have different gene_names associated with it, without creating a unique gene_id for every transcript_id? Thank you!