-
-
Save muammar/6d298f7fa5700efdbd01ae697f63e089 to your computer and use it in GitHub Desktop.
Example Error of a Neural Network trained for regression with Pytorch
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| import time | |
| import datetime | |
| import torch | |
| from mlchemistry.data.visualization import parity | |
| torch.set_printoptions(precision=10) | |
| class NeuralNetwork(torch.nn.Module): | |
| """Neural Network Regression with Pytorch | |
| Parameters | |
| ---------- | |
| hiddenlayers : tuple | |
| Structure of hidden layers in the neural network. | |
| activation : str | |
| The activation function. | |
| """ | |
| def __init__(self, hiddenlayers=(3, 3), activation='relu'): | |
| super(NeuralNetwork, self).__init__() | |
| self.hiddenlayers = hiddenlayers | |
| self.activation = activation | |
| def prepare_model(self, input_dimension, data=None): | |
| """Prepare the model | |
| Parameters | |
| ---------- | |
| input_dimension : int | |
| Input's dimension. | |
| data : object | |
| DataSet object created from the handler. | |
| """ | |
| activation = {'tanh': torch.nn.Tanh, 'relu': torch.nn.ReLU, | |
| 'celu': torch.nn.CELU} | |
| print() | |
| print('Model Training') | |
| print('==============') | |
| print('Number of hidden-layers: {}' .format(len(self.hiddenlayers))) | |
| print('Structure of Neural Net: {}' . | |
| format('(input, ' + str(self.hiddenlayers)[1:-1] + ', output)')) | |
| layers = range(len(self.hiddenlayers) + 1) | |
| unique_element_symbols = data.unique_element_symbols['trainingset'] | |
| symbol_model_pair = [] | |
| self.output_layer_index = {} | |
| for symbol in unique_element_symbols: | |
| linears = [] | |
| intercept = (data.max_energy + data.min_energy) / 2. | |
| intercept = torch.nn.Parameter(torch.tensor(intercept, | |
| requires_grad=True)) | |
| slope = (data.max_energy - data.min_energy) / 2. | |
| slope = torch.nn.Parameter(torch.tensor(slope, requires_grad=True)) | |
| print(intercept, slope) | |
| intercept_name = 'intercept_' + symbol | |
| slope_name = 'slope_' + symbol | |
| self.register_parameter(intercept_name, intercept) | |
| self.register_parameter(slope_name, slope) | |
| for index in layers: | |
| # This is the input layer | |
| if index == 0: | |
| out_dimension = self.hiddenlayers[0] | |
| _linear = torch.nn.Linear(input_dimension, | |
| out_dimension) | |
| linears.append(_linear) | |
| linears.append(activation[self.activation]()) | |
| # This is the output layer | |
| elif index == len(self.hiddenlayers): | |
| inp_dimension = self.hiddenlayers[index - 1] | |
| out_dimension = 1 | |
| self.output_layer_index[symbol] = index | |
| _linear = torch.nn.Linear(inp_dimension, out_dimension) | |
| linears.append(_linear) | |
| # These are hidden-layers | |
| else: | |
| inp_dimension = self.hiddenlayers[index - 1] | |
| out_dimension = self.hiddenlayers[index] | |
| _linear = torch.nn.Linear(inp_dimension, out_dimension) | |
| linears.append(_linear) | |
| linears.append(activation[self.activation]()) | |
| # Stacking up the layers. | |
| linears = torch.nn.Sequential(*linears) | |
| symbol_model_pair.append([symbol, linears]) | |
| self.linears = torch.nn.ModuleDict(symbol_model_pair) | |
| print(self.linears) | |
| # Iterate over all modules and just intialize those that are a linear | |
| # layer. | |
| for m in self.modules(): | |
| if isinstance(m, torch.nn.Linear): | |
| # nn.init.normal_(m.weight) # , mean=0, std=0.01) | |
| torch.nn.init.xavier_uniform_(m.weight) | |
| def forward(self, X): | |
| """Forward propagation | |
| This is forward propagation and it returns the atomic energy. | |
| Parameters | |
| ---------- | |
| X : list | |
| List of inputs in the feature space. | |
| Returns | |
| ------- | |
| outputs : tensor | |
| A list of tensors with energies per image. | |
| """ | |
| outputs = [] | |
| for hash in X: | |
| image = X[hash] | |
| atomic_energies = [] | |
| for symbol, x in image: | |
| x = self.linears[symbol](x) | |
| intercept_name = 'intercept_' + symbol | |
| slope_name = 'slope_' + symbol | |
| for name, param in self.named_parameters(): | |
| if intercept_name == name: | |
| intercept = param | |
| elif slope_name == name: | |
| slope = param | |
| x = (slope * x) + intercept | |
| atomic_energies.append(x) | |
| atomic_energies = torch.cat(atomic_energies) | |
| image_energy = torch.sum(atomic_energies) | |
| outputs.append(image_energy) | |
| outputs = torch.stack(outputs) | |
| return outputs | |
| def train(self, inputs, targets, model=None, data=None, optimizer=None, | |
| lr=None, weight_decay=None, regularization=None, epochs=100, | |
| convergence=None, lossfxn=None): | |
| """Train the model | |
| Parameters | |
| ---------- | |
| inputs : dict | |
| Dictionary with hashed feature space. | |
| epochs : int | |
| Number of full training cycles. | |
| targets : list | |
| The expected values that the model has to learn aka y. | |
| model : object | |
| The NeuralNetwork class. | |
| data : object | |
| DataSet object created from the handler. | |
| lr : float | |
| Learning rate. | |
| weight_decay : float | |
| Weight decay passed to the optimizer. Default is 0. | |
| regularization : float | |
| This is the L2 regularization. It is not the same as weight decay. | |
| convergence : dict | |
| Instead of using epochs, users can set a convergence criterion. | |
| lossfxn : obj | |
| A loss function object. | |
| """ | |
| #old_state_dict = {} | |
| #for key in model.state_dict(): | |
| # old_state_dict[key] = model.state_dict()[key].clone() | |
| targets = torch.tensor(targets, requires_grad=False) | |
| # Define optimizer | |
| if optimizer is None: | |
| optimizer = torch.optim.Adam(model.parameters(), lr=lr, | |
| weight_decay=weight_decay) | |
| print() | |
| print('{:6s} {:19s} {:8s}'.format('Epoch', 'Time Stamp', 'Loss')) | |
| print('{:6s} {:19s} {:8s}'.format('------', | |
| '-------------------', '---------')) | |
| initial_time = time.time() | |
| _loss = [] | |
| _rmse = [] | |
| epoch = 0 | |
| while True: | |
| epoch += 1 | |
| outputs = model(inputs) | |
| if lossfxn is None: | |
| loss, rmse = self.loss_function(outputs, targets, optimizer, data) | |
| else: | |
| raise('I do not know what to do') | |
| _loss.append(loss) | |
| _rmse.append(rmse) | |
| ts = time.time() | |
| ts = datetime.datetime.fromtimestamp(ts).strftime('%Y-%m-%d ' | |
| '%H:%M:%S') | |
| print('{:6d} {} {:8e} {:8f}' .format(epoch, ts, loss, rmse)) | |
| if convergence is None and epoch == epochs: | |
| break | |
| elif (convergence is not None and rmse < convergence['energy']): | |
| break | |
| training_time = time.time() - initial_time | |
| print('Training the model took {}...' .format(training_time)) | |
| print('outputs') | |
| print(outputs) | |
| print('targets') | |
| print(targets) | |
| import matplotlib.pyplot as plt | |
| plt.plot(list(range(epoch)), _loss, label='loss') | |
| plt.plot(list(range(epoch)), _rmse, label='rmse/atom') | |
| plt.legend(loc='upper left') | |
| plt.show() | |
| parity(outputs.detach().numpy(), targets.detach().numpy()) | |
| #new_state_dict = {} | |
| #for key in model.state_dict(): | |
| # new_state_dict[key] = model.state_dict()[key].clone() | |
| #for key in old_state_dict: | |
| # if not (old_state_dict[key] == new_state_dict[key]).all(): | |
| # print('Diff in {}'.format(key)) | |
| # else: | |
| # print('No diff in {}'.format(key)) | |
| #print() | |
| #for symbol in data.unique_element_symbols['trainingset']: | |
| # model = model.linears[symbol] | |
| # print('Optimized parameters for {} symbol' .format(symbol)) | |
| # for index, param in enumerate(model.parameters()): | |
| # print('Index {}' .format(index)) | |
| # print(param) | |
| # try: | |
| # print('Gradient', param.grad.sum()) | |
| # except AttributeError: | |
| # print('No gradient?') | |
| # print() | |
| def loss_function(self, outputs, targets, optimizer, data): | |
| """Default loss function | |
| If user does not input loss function we provide mean-squared error loss | |
| function. | |
| Parameters | |
| ---------- | |
| outputs : tensor | |
| Outputs of the model. | |
| targets : tensor | |
| Expected value of outputs. | |
| optimizer : obj | |
| An optimizer object to minimize the loss function error. | |
| data : obj | |
| A data object from mlchem. | |
| Returns | |
| ------- | |
| loss : tensor | |
| The value of the loss function. | |
| rmse : float | |
| Value of the root-mean squared error per atom. | |
| """ | |
| optimizer.zero_grad() # clear previous gradients | |
| criterion = torch.nn.MSELoss(reduction='sum') | |
| atoms_per_image = torch.tensor(data.atoms_per_image, | |
| requires_grad=False, | |
| dtype=torch.float) | |
| outputs_atom = torch.div(outputs, atoms_per_image) | |
| targets_atom = torch.div(targets, atoms_per_image) | |
| loss = criterion(outputs_atom, targets_atom) * .5 | |
| loss.backward() | |
| optimizer.step() | |
| rmse = torch.sqrt(loss).item() | |
| return loss, rmse |
Sign up for free
to join this conversation on GitHub.
Already have an account?
Sign in to comment
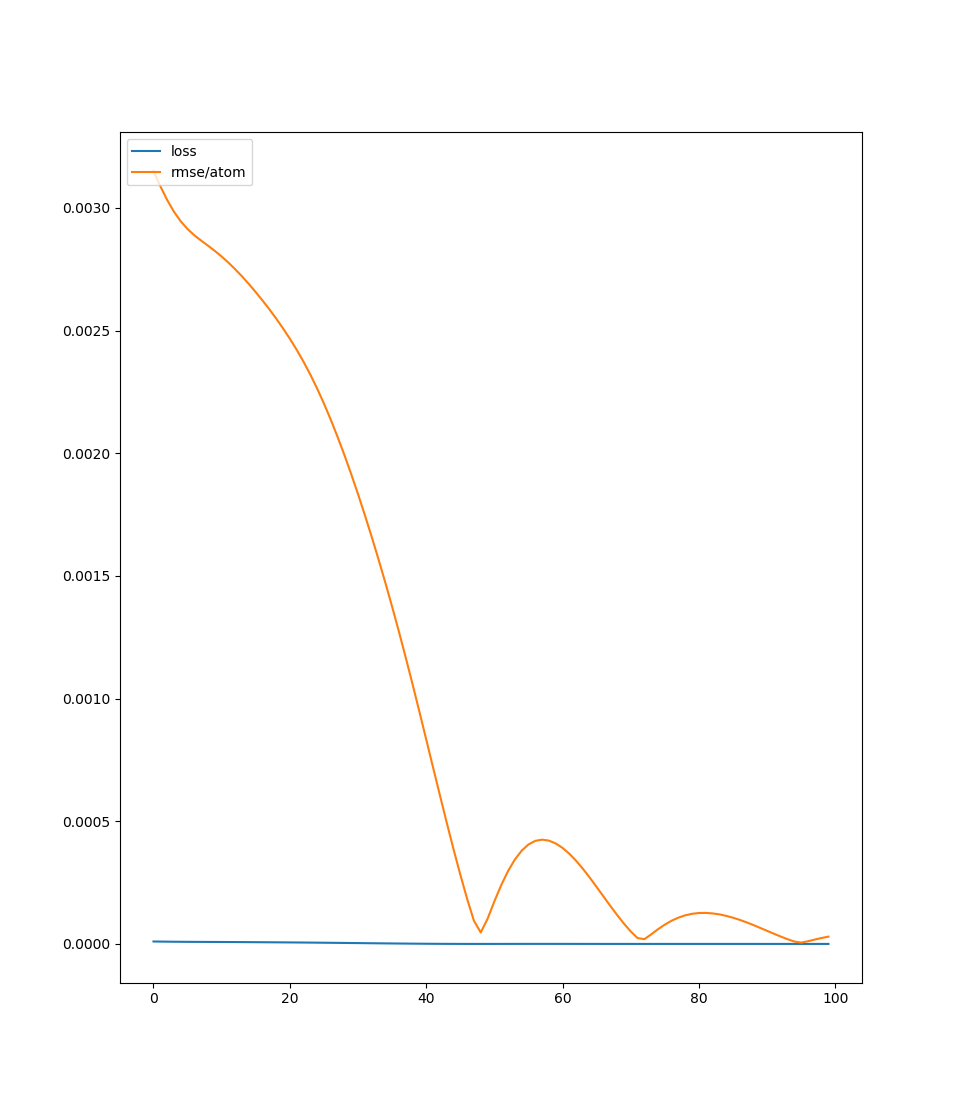

Output of running script above