Last active
June 10, 2018 16:29
-
-
Save quantum-kite/b2457db46dbff9ad02a56443255ace46 to your computer and use it in GitHub Desktop.
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| """ Bond disorder | |
| Lattice : Honeycomb 1[nm] interatomic distance and t=1[eV] hopping; | |
| Disorder : StructuralDisorder class bond and vacancy disorder; | |
| Configuration : size of the system 512x512, without domain decomposition (nx=ny=1), periodic boundary conditions, | |
| double precision, manual scaling; | |
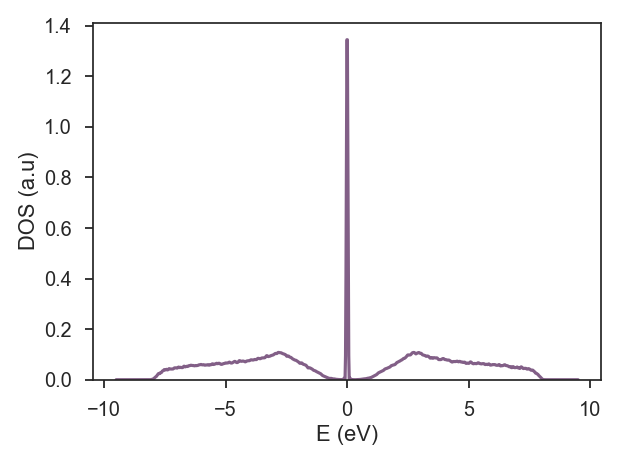
| Calculation : dos; | |
| Modification : magnetic field is off; | |
| """ | |
| import numpy as np | |
| import pybinding as pb | |
| import kite | |
| def honeycomb_lattice(onsite=(0, 0)): | |
| """Make a honeycomb lattice with nearest neighbor hopping""" | |
| theta = np.pi / 3 | |
| a1 = np.array([1 + np.cos(theta), np.sin(theta)]) | |
| a2 = np.array([0, 2 * np.sin(theta)]) | |
| # create a lattice with 2 primitive vectors | |
| lat = pb.Lattice( | |
| a1=a1, | |
| a2=a2 | |
| ) | |
| # Add sublattices | |
| lat.add_sublattices( | |
| # name, position, and onsite potential | |
| ('A', [0, 0], onsite[0]), | |
| ('B', [1, 0], onsite[1]) | |
| ) | |
| # Add hoppings | |
| lat.add_hoppings( | |
| # inside the main cell, between which atoms, and the value | |
| ([0, 0], 'A', 'B', - 1), | |
| # between neighboring cells, between which atoms, and the value | |
| ([-1, 0], 'A', 'B', - 1), | |
| ([-1, 1], 'A', 'B', - 1), | |
| ) | |
| # Add bond disorder as an object of a class StructuralDisorder. In this manner we can add onsite and bond defects | |
| # with a specific concentration, which will be added to the simulated system. The procedure for adding is same | |
| # as adding the hopping, with the difference that the bond disorded is not bounded to one site in the [0, 0] | |
| # unit cell. | |
| node0 = [[+0, +0], 'A'] | |
| node1 = [[+0, +0], 'B'] | |
| node2 = [[+1, +0], 'A'] | |
| node3 = [[+0, +1], 'B'] | |
| node4 = [[+0, +1], 'A'] | |
| node5 = [[-1, +1], 'B'] | |
| struc_disorder_one = kite.StructuralDisorder(lat, concentration=0.05) | |
| struc_disorder_one.add_structural_disorder( | |
| # add bond disorder in the form [from unit cell], 'sublattice_from', [to_unit_cell], 'sublattice_to', value: | |
| (*node0, *node1, 1), | |
| (*node1, *node2, 1), | |
| (*node2, *node3, 1), | |
| (*node3, *node4, 1), | |
| (*node4, *node5, 1), | |
| (*node5, *node0, 1), | |
| # in this way we can add onsite disorder in the form [unit cell], 'sublattice', value | |
| ([+0, +0], 'B', 0.3) | |
| ) | |
| # It is possible to add multiple different disorder type which should be forwarded to the export_lattice function | |
| # as a list. | |
| struc_disorder_two = kite.StructuralDisorder(lat, concentration=0.2) | |
| struc_disorder_two.add_structural_disorder( | |
| (*node0, *node1, 0.4), | |
| (*node4, *node5, 0.4), | |
| (*node5, *node0, 0.4), | |
| ([+0, +0], 'B', 0.4) | |
| ) | |
| struc_disorder_two.add_vacancy('B') | |
| struc_disorder_three = kite.StructuralDisorder(lat, concentration=0.01) | |
| struc_disorder_three.add_vacancy('A') | |
| # if there is disorder it should be returned separately from the lattice | |
| return lat, [struc_disorder_one, struc_disorder_two, struc_disorder_three] | |
| # load a honeycomb lattice and structural_disorder | |
| lattice, disorder_structural = honeycomb_lattice() | |
| # number of decomposition parts in each direction of matrix. | |
| # This divides the lattice into various sections, each of which is calculated in parallel | |
| nx = ny = 1 | |
| # number of unit cells in each direction. | |
| lx = ly = 512 | |
| # make config object which caries info about | |
| # - the number of decomposition parts [nx, ny], | |
| # - lengths of structure [lx, ly] | |
| # - boundary conditions, setting True as periodic boundary conditions, and False elsewise, | |
| # - info if the exported hopping and onsite data should be complex, | |
| # - info of the precision of the exported hopping and onsite data, 0 - float, 1 - double, and 2 - long double. | |
| # - scaling, if None it's automatic, if present select spectrum_bound=[e_min, e_max] | |
| configuration = kite.Configuration(divisions=[nx, ny], length=[lx, ly], boundaries=[True, True], | |
| is_complex=False, precision=1, spectrum_range=[-15, 15]) | |
| # require the calculation of DOS | |
| calculation = kite.Calculation(configuration) | |
| calculation.dos(num_moments=1024, num_random=1, num_disorder=1, num_points=1000) | |
| # configure the *.h5 file | |
| kite.config_system(lattice, configuration, calculation, filename='structural_disorder.h5', | |
| disorder_structural=disorder_structural) |
Author
quantum-kite
commented
Jun 5, 2018
Sign up for free
to join this conversation on GitHub.
Already have an account?
Sign in to comment