Last active
August 6, 2021 11:07
-
-
Save NatureGeorge/62cdb644154fb11f3fa73f02dbcf2b9b to your computer and use it in GitHub Desktop.
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Copyright (C) 2002, Thomas Hamelryck (thamelry@binf.ku.dk) | |
| # | |
| # This file is part of the Biopython distribution and governed by your | |
| # choice of the "Biopython License Agreement" or the "BSD 3-Clause License". | |
| # Please see the LICENSE file that should have been included as part of this | |
| # package. | |
| """Turn an mmCIF file into a dictionary.""" | |
| from Bio.File import as_handle | |
| class MMCIF2Dict(dict): | |
| """Parse a mmCIF file and return a dictionary.""" | |
| def __init__(self, filename): | |
| """Parse a mmCIF file and return a dictionary. | |
| Arguments: | |
| - file - name of the PDB file OR an open filehandle | |
| """ | |
| self.quote_chars = ["'", '"'] | |
| self.whitespace_chars = [" ", "\t"] | |
| with as_handle(filename) as handle: | |
| loop_flag = False | |
| key = None | |
| tokens = self._tokenize(handle) | |
| try: | |
| token = next(tokens) | |
| except StopIteration: | |
| return # for Python 3.7 and PEP 479 | |
| self[token[0:5]] = token[5:] | |
| i = 0 | |
| n = 0 | |
| for token in tokens: | |
| if token.lower() == "loop_": | |
| loop_flag = True | |
| keys = [] | |
| i = 0 | |
| n = 0 | |
| continue | |
| elif loop_flag: | |
| # The second condition checks we are in the first column | |
| # Some mmCIF files (e.g. 4q9r) have values in later columns | |
| # starting with an underscore and we don't want to read | |
| # these as keys | |
| if token.startswith("_") and (n == 0 or i % n == 0): | |
| if i > 0: | |
| loop_flag = False | |
| else: | |
| self[token] = [] | |
| keys.append(token) | |
| n += 1 | |
| continue | |
| else: | |
| self[keys[i % n]].append(token) | |
| i += 1 | |
| continue | |
| if key is None: | |
| key = token | |
| else: | |
| self[key] = [token] | |
| key = None | |
| # Private methods | |
| def _splitline(self, line): | |
| # See https://www.iucr.org/resources/cif/spec/version1.1/cifsyntax for the syntax | |
| in_token = False | |
| # quote character of the currently open quote, or None if no quote open | |
| quote_open_char = None | |
| start_i = 0 | |
| for (i, c) in enumerate(line): | |
| if c in self.whitespace_chars: | |
| if in_token and not quote_open_char: | |
| in_token = False | |
| yield line[start_i:i] | |
| elif c in self.quote_chars: | |
| if not quote_open_char and not in_token: | |
| quote_open_char = c | |
| in_token = True | |
| start_i = i + 1 | |
| elif c == quote_open_char and ( | |
| i + 1 == len(line) or line[i + 1] in self.whitespace_chars | |
| ): | |
| quote_open_char = None | |
| in_token = False | |
| yield line[start_i:i] | |
| elif c == "#" and not in_token: | |
| # Skip comments. "#" is a valid non-comment char inside of a | |
| # quote and inside of an unquoted token (!?!?), so we need to | |
| # check that the current char is not in a token. | |
| return | |
| elif not in_token: | |
| in_token = True | |
| start_i = i | |
| if in_token: | |
| yield line[start_i:] | |
| if quote_open_char: | |
| raise ValueError("Line ended with quote open: " + line) | |
| def _tokenize(self, handle): | |
| empty = True | |
| for line in handle: | |
| empty = False | |
| if line.startswith("#"): | |
| continue | |
| elif line.startswith(";"): | |
| # The spec says that leading whitespace on each line must be | |
| # preserved while trailing whitespace may be stripped. The | |
| # trailing newline must be stripped. | |
| token_buffer = [line[1:].rstrip()] | |
| for line in handle: | |
| line = line.rstrip() | |
| if line.startswith(";"): | |
| yield "\n".join(token_buffer) | |
| line = line[1:] | |
| if line and not line[0] in self.whitespace_chars: | |
| raise ValueError("Missing whitespace") | |
| break | |
| token_buffer.append(line) | |
| else: | |
| raise ValueError("Missing closing semicolon") | |
| yield from self._splitline(line.strip()) | |
| if empty: | |
| raise ValueError("Empty file.") |
Identifiers Related
- biojava/biojava#220: Better support for symmetry in the Structure model
- biojava/biojava#469: Add better support for asym chain ids as well as auth ids
- biojava/biojava#479: Refactoring of structure data model
- biojava/biojava#801: Biological assembly expansion: chain ids should contain both operator ids in binary expression case
- biojava/biojava#802: Assembly chain ids for cases with composed operators in assembly expansion
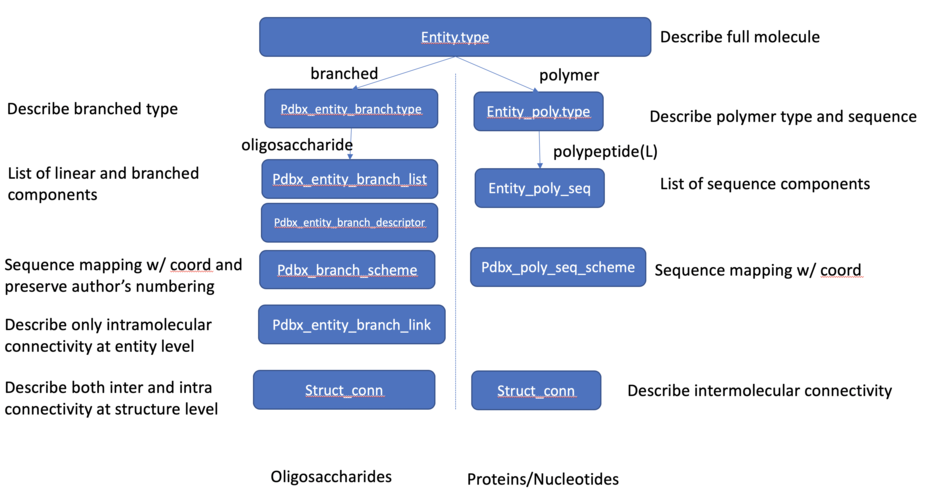
Carbohydrate Related
- biojava/biojava#868: Minimal read support for files with 'branched' entities
TODO (2020-09-11)
Plan to implement lark-parser to rewrite MMCIF2DictPlus.
Sign up for free
to join this conversation on GitHub.
Already have an account?
Sign in to comment
Usage
Output
2Q4N{"_pdbx_struct_assembly_gen.assembly_id": ["1"], "_pdbx_struct_assembly_gen.asym_id_list": ["A,B"], "_pdbx_struct_assembly_gen.oper_expression": ["1,2"], "_pdbx_struct_oper_list.id": ["1", "2"], "_pdbx_struct_oper_list.symmetry_operation": ["x,y,z", "-x,-y+2,z"], "data_": "2Q4N"}3HL2{"_pdbx_struct_assembly_gen.assembly_id": ["1", "1", "2", "2"], "_pdbx_struct_assembly_gen.asym_id_list": ["A,B,F,G,H,I,J,K,Q,R", "A,B,E,F,G,H,I,J,K,Q,R,U", "C,D,L,M,N,O,P,S,T", "C,D,E,L,M,N,O,P,S,T,U"], "_pdbx_struct_assembly_gen.oper_expression": ["1", "2", "3", "2"], "_pdbx_struct_oper_list.id": ["1", "2", "3"], "_pdbx_struct_oper_list.symmetry_operation": ["x,x-y-1,-z", "x,y,z", "-x+y,y,-z+1/3"], "data_": "3HL2"}1M11{ "data_": "1M11", "_pdbx_struct_assembly_gen.asym_id_list": ["A,B,C,D", "A,B,C,D", "A,B,C,D", "A,B,C,D", "A,B,C,D"], "_pdbx_struct_assembly_gen.assembly_id": ["1", "2", "3", "4", "5"], "_pdbx_struct_assembly_gen.oper_expression": ["(1-60)", "1", "(1-5)", "(1,2,6,10,23,24)", "P"], "_pdbx_struct_oper_list.id": ["P", "1", "2", "3", "4", "5", "6", "7", "8", "9", "10", "11", "12", "13", "14", "15", "16", "17", "18", "19", "20", "21", "22", "23", "24", "25", "26", "27", "28", "29", "30", "31", "32", "33", "34", "35", "36", "37", "38", "39", "40", "41", "42", "43", "44", "45", "46", "47", "48", "49", "50", "51", "52", "53", "54", "55", "56", "57", "58", "59", "60"], "_pdbx_struct_oper_list.symmetry_operation": ["?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?"] }1M4X{ "data_": "1M4X", "_pdbx_struct_assembly_gen.asym_id_list": ["A,B,C", "A,B,C", "A,B,C", "A,B,C", "A,B,C", "A,B,C", "A,B,C"], "_pdbx_struct_assembly_gen.assembly_id": ["1", "2", "3", "4", "5", "6", "7"], "_pdbx_struct_assembly_gen.oper_expression": ["(1-60)(61-88)", "(61-88)", "(1-5)(61-88)", "(1,2,6,10,23,24)(61-88)", "(1-5)(63-68)", "(1,10,23)(61,62,69-88)", "(P)(61-88)"], "_pdbx_struct_oper_list.id": ["P", "1", "2", "3", "4", "5", "6", "7", "8", "9", "10", "11", "12", "13", "14", "15", "16", "17", "18", "19", "20", "21", "22", "23", "24", "25", "26", "27", "28", "29", "30", "31", "32", "33", "34", "35", "36", "37", "38", "39", "40", "41", "42", "43", "44", "45", "46", "47", "48", "49", "50", "51", "52", "53", "54", "55", "56", "57", "58", "59", "60", "61", "62", "63", "64", "65", "66", "67", "68", "69", "70", "71", "72", "73", "74", "75", "76", "77", "78", "79", "80", "81", "82", "83", "84", "85", "86", "87", "88"], "_pdbx_struct_oper_list.symmetry_operation": ["?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?", "?"] }